Computational Analysis for Cancer Pathology
Histopathology interpretation is expert-driven, inherently complex, and difficult to scale. Tsebix develops computational methods to assist pathologists by making tissue analysis more consistent, accessible, and quantifiable.
- Calibrated cancer risk score
- Cloud-based analysis
- Whole-slide image support
- Microscope image compatibility
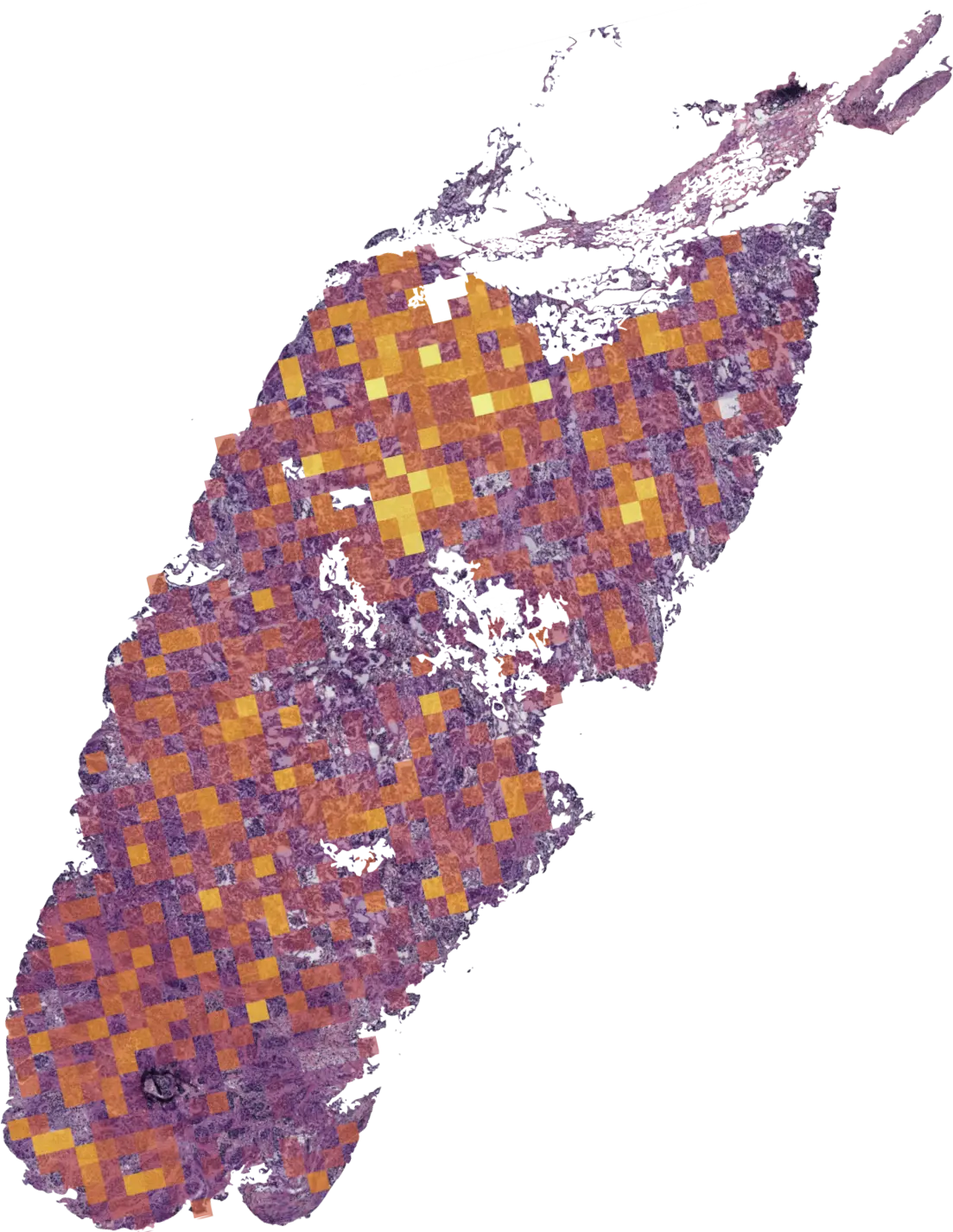
- Interpretable heatmaps
- AI-assisted morphological description
Clinical Context
Pathology sits at the center of cancer care, yet many regions face delays, diagnostic variability, and limited access to specialist expertise. This gap is widening as global cancer incidence continues to rise.
Rising Demand
Global cancer incidence is increasing, placing growing pressure on diagnostic systems and pathology services.
The Expertise Gap
Many healthcare systems face a shortage of trained pathologists, contributing to delays in critical diagnostic workflows.
Universal Access
Digital pathology infrastructure varies widely across institutions. Some rely on high-end slide scanners, while others operate with standard optical microscopes and digital cameras.
Tsebix is developing computational tools that help pathologists analyze histopathology images more efficiently across different workflows, settings, and resource levels. The goal is not to replace clinical expertise, but to augment it where it is most needed.
Our work prioritizes reliability, accessibility, and trust.
Platform
The Tsebix platform provides a browser-based environment for analyzing histopathology images and reviewing model outputs.
Current Status: Research Use Only (RUO) — The platform is live and accessible to research collaborators. Models have been trained on TCGA and independently validated on CPTAC across eight cancer types. Clinical validation activities are underway with hospital partners. The platform is under active development; additional cancer types are currently being incorporated.
Inputs
Whole Slide Images
Standard digital pathology formats.
Microscope Images
Images captured directly
from optical microscopes without dedicated slide scanners.
Outputs
Cancer Risk Score
A calibrated probability estimate indicating whether the
tissue contains cancer-related morphological features.
Heatmaps
Spatial visualization of regions
contributing to the model’s prediction. These maps guide pathologists to areas of
interest.
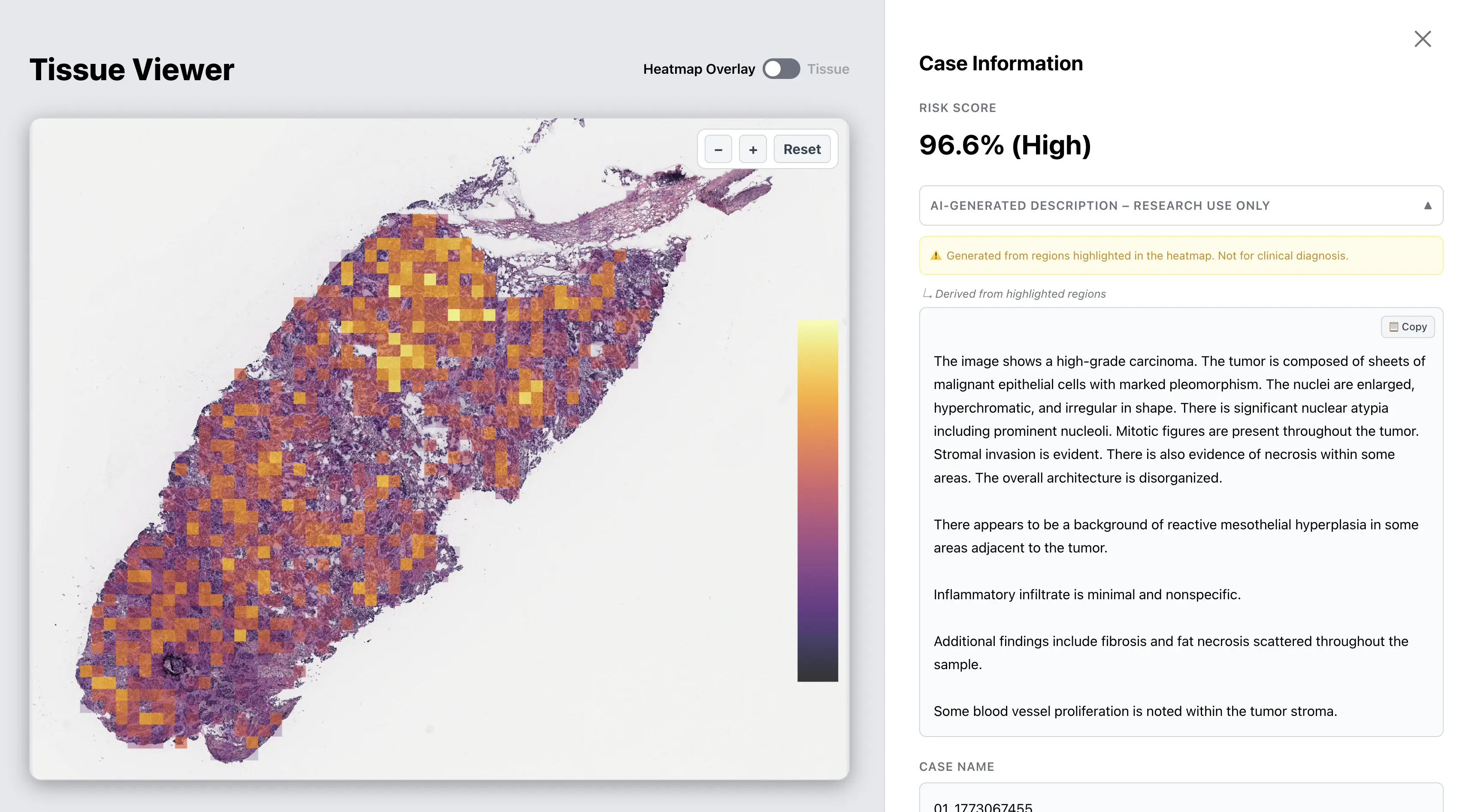
AI-Generated Description
The system can
generate descriptive summaries of morphological features from image regions highlighted
by the model.
Case list view — processed cases with risk scores
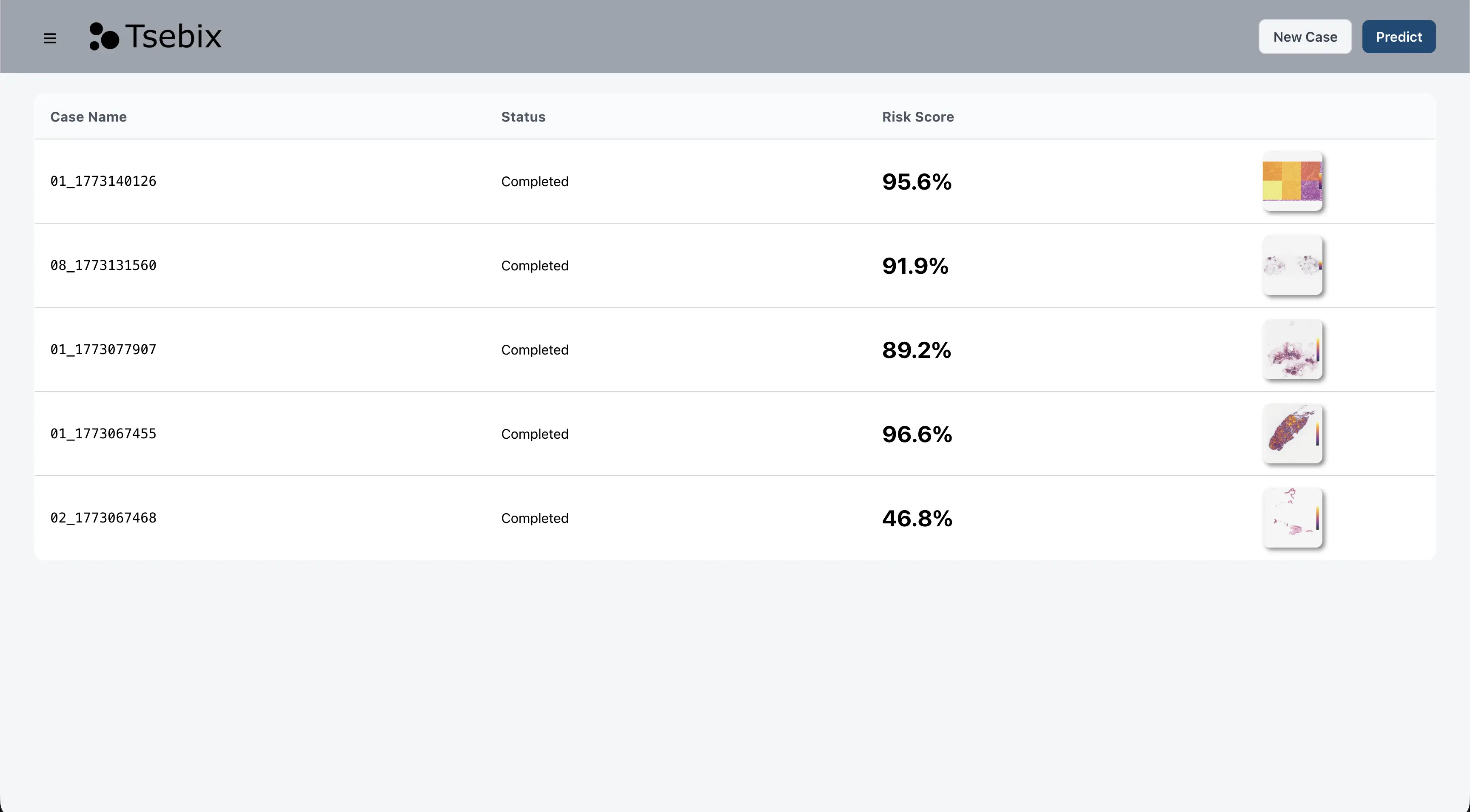
Platform interface — showing analysis results for a sample histopathology case. Data shown is from a public research dataset.
Technology
Whole-slide histopathology images contain billions of pixels, while diagnostic labels are typically available only at the slide level. The Tsebix pipeline analyzes these large images to identify regions most relevant to the model’s prediction without requiring pixel-level annotations. This approach enables efficient analysis of large tissue images while producing calibrated probability estimates and spatial heatmaps that help guide interpretation.
Image Representation Models
Foundation models used to encode histopathology image features.
Attention-Based Learning
Models designed to identify relevant regions within large tissue images.
Calibration
Methods used to ensure that model outputs correspond to reliable probability estimates.
Multimodal Models
Vision-language models used to generate descriptive summaries of image regions.
Infrastructure
Cloud-based architecture allowing large-image processing through a browser interface.
Validation
Training & Evaluation
Models were trained using The Cancer Genome Atlas (TCGA) and evaluated on independent data from the Clinical Proteomic Tumor Analysis Consortium (CPTAC). The current pan-cancer model is evaluated on a binary classification task (cancer vs. normal tissue) across multiple cancer types.
Tissue Types
Evaluated across multiple cancer tissues including:
- Breast
- Lung
- Colorectal
- Renal
- Ovarian
- Head and Neck
- Endometrial
Performance Summary
Research
Tsebix focuses on computational pathology methods for cancer analysis, including interpretable AI models and multimodal medical AI.
Current Work
Our ongoing efforts are focused on:
- Expansion of model validation with clinical collaborators
- Evaluation with practicing pathologists in real workflows
- Extension of the platform to additional cancer tissue types
- Continued refinement of computational pathology models
- Development of models exploring the prediction of molecular biomarkers from histopathology images
Vision
Tsebix aims to develop reliable computational tools that assist pathologists in the analysis of
histopathology images and expand access to advanced digital pathology technologies.
The platform is designed to augment clinical expertise rather than replace it.
Team
Jaime F. Delgado Saa, Ph.D. — Founder
Jaime holds a Ph.D. in Electronics Engineering and conducted postdoctoral research at the University of Geneva, Faculty of Medicine, where his work focused on machine learning, probabilistic graphical models, and the analysis of biological signals. He founded Tsebix to apply rigorous computational methods to histopathology, with the goal of building tools that function reliably in real clinical settings.
LinkedInFrequently Asked Questions
Is Tsebix intended for clinical diagnosis?
The platform is currently intended for research and evaluation purposes. Model outputs require expert interpretation and are not intended for clinical decision-making.
What image formats are supported?
The platform supports standard digital pathology whole-slide image formats as well as microscope images captured using optical microscopes and digital cameras.
What data were used to train and evaluate the models?
Models were trained using data from The Cancer Genome Atlas (TCGA) and evaluated on independent datasets from the Clinical Proteomic Tumor Analysis Consortium (CPTAC).
Who can request access to the platform?
Access is currently available to academic researchers and clinical collaborators interested in evaluating computational pathology tools.
What types of cancer are currently supported?
Current models have been evaluated across multiple tissue types including breast, lung, colorectal, renal, ovarian, head and neck, and endometrial cancers.